Supervisor: Detlev Arendt (EMBL Heidelberg), Co-supervisor Gert Wörheide (LMU Munich)
Student: Fabian Ruperti
Objectives: The nature of the last common ancestor of the Metazoa is largely unsolved, due to uncertainties in the branching pattern of non-bilaterian clades and, consequently, in the significance of the apparent morphological simplicity. For instance, our understanding and interpretation of the simple body plan of sponges and placozoa largely depends on their actual phylogenetic position in the animal tree of life. One distinct feature of both phyla is the absence of neurons and muscles which may be due to an early branch-off and suggests them to be primarily simple; should they be closer related to cnidarians than for instance ctenophores, this could account for a secondary simplification or – more probably – an independent evolution of a nervous system in ctenophores. However, despite the absence of a nervous system, the genome and transcriptome of sponges and placozoans show the occurrence of typical synaptic proteins. Also, when treated with neurotransmitters and neuropeptides common in other metazoans they show behavioral reactions suggesting a primeval neuron-like signal transduction. The PhD student of project 4 will focus on the protein specific interactions leading to the uptake and transduction of “neuronal” signals in sponges and placozoans by means of proteomics.
O1:Establish protocols for cultivation and treatment of sponges and placozoans and subsequent preparation for mass spectrometry.
O2: Conduct thermal protein profiling (TPP) proteomics experiments on in vivo treated sponges and placozoans and use established bioinformatic pipelines to infer candidate proteins and protein complexes.
O3: Compare candidate proteins and protein complexes to known neuronal protein modules. Support findings by biophysical characterization of purified proteins, in situ immunostaining and Co-Immunoprecipitation.
Secondments: Grace McCormack (Galway), Gert Wörheide (Munich), Lyubomir Penev (Pensoft Publishers, Sofia).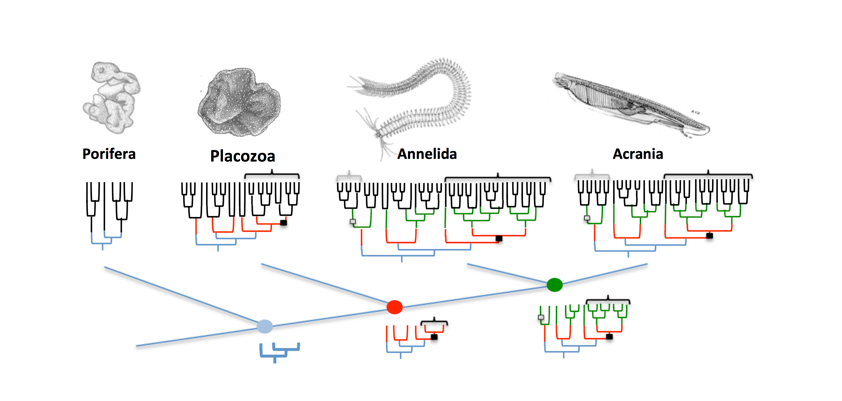
Student: Fabian Ruperti
Objectives: The nature of the last common ancestor of the Metazoa is largely unsolved, due to uncertainties in the branching pattern of non-bilaterian clades and, consequently, in the significance of the apparent morphological simplicity. For instance, our understanding and interpretation of the simple body plan of sponges and placozoa largely depends on their actual phylogenetic position in the animal tree of life. One distinct feature of both phyla is the absence of neurons and muscles which may be due to an early branch-off and suggests them to be primarily simple; should they be closer related to cnidarians than for instance ctenophores, this could account for a secondary simplification or – more probably – an independent evolution of a nervous system in ctenophores. However, despite the absence of a nervous system, the genome and transcriptome of sponges and placozoans show the occurrence of typical synaptic proteins. Also, when treated with neurotransmitters and neuropeptides common in other metazoans they show behavioral reactions suggesting a primeval neuron-like signal transduction. The PhD student of project 4 will focus on the protein specific interactions leading to the uptake and transduction of “neuronal” signals in sponges and placozoans by means of proteomics.
O1:Establish protocols for cultivation and treatment of sponges and placozoans and subsequent preparation for mass spectrometry.
O2: Conduct thermal protein profiling (TPP) proteomics experiments on in vivo treated sponges and placozoans and use established bioinformatic pipelines to infer candidate proteins and protein complexes.
O3: Compare candidate proteins and protein complexes to known neuronal protein modules. Support findings by biophysical characterization of purified proteins, in situ immunostaining and Co-Immunoprecipitation.
Secondments: Grace McCormack (Galway), Gert Wörheide (Munich), Lyubomir Penev (Pensoft Publishers, Sofia).
