In this study, the complete mitogenome of an invasive pest slug, Arion vulgaris Moquin-Tandon, 1855 (Stylommatophora: Arionidae), was sequenced using next generation sequencing, analysed and compared with other stylommatophorans. The mitogenome of A. vulgaris measures 14,547 bp and contains 13 protein-coding, two rRNA, 22 tRNA genes, and one control region, with an A + T content of 70.20%. All protein coding genes (PCGs) are initiated with ATN codons except for COX1, ND5 and ATP8 and all are ended with TAR or T-stop codons. All tRNAs were folded into a clover-leaf secondary structure except for trnC and trnS1 (AGN). Phylogenetic analyses confirmed the position of A. vulgaris within the superfamily Arionoidea, recovered a sister group relationship between Arionoidea and Orthalicoidea, and supported monophyly of all currently recognized superfamilies within Stylommatophora except for the superfamily Helicoidea. Initial diversification time of the Stylommatophora was estimated as 138.55 million years ago corresponding to Early Cretaceous. The divergence time of A. vulgaris and Arion rufus (Linnaeus, 1758) was estimated as 15.24 million years ago corresponding to one of Earth’s most recent, global warming events, the Mid-Miocene Climatic Optimum.
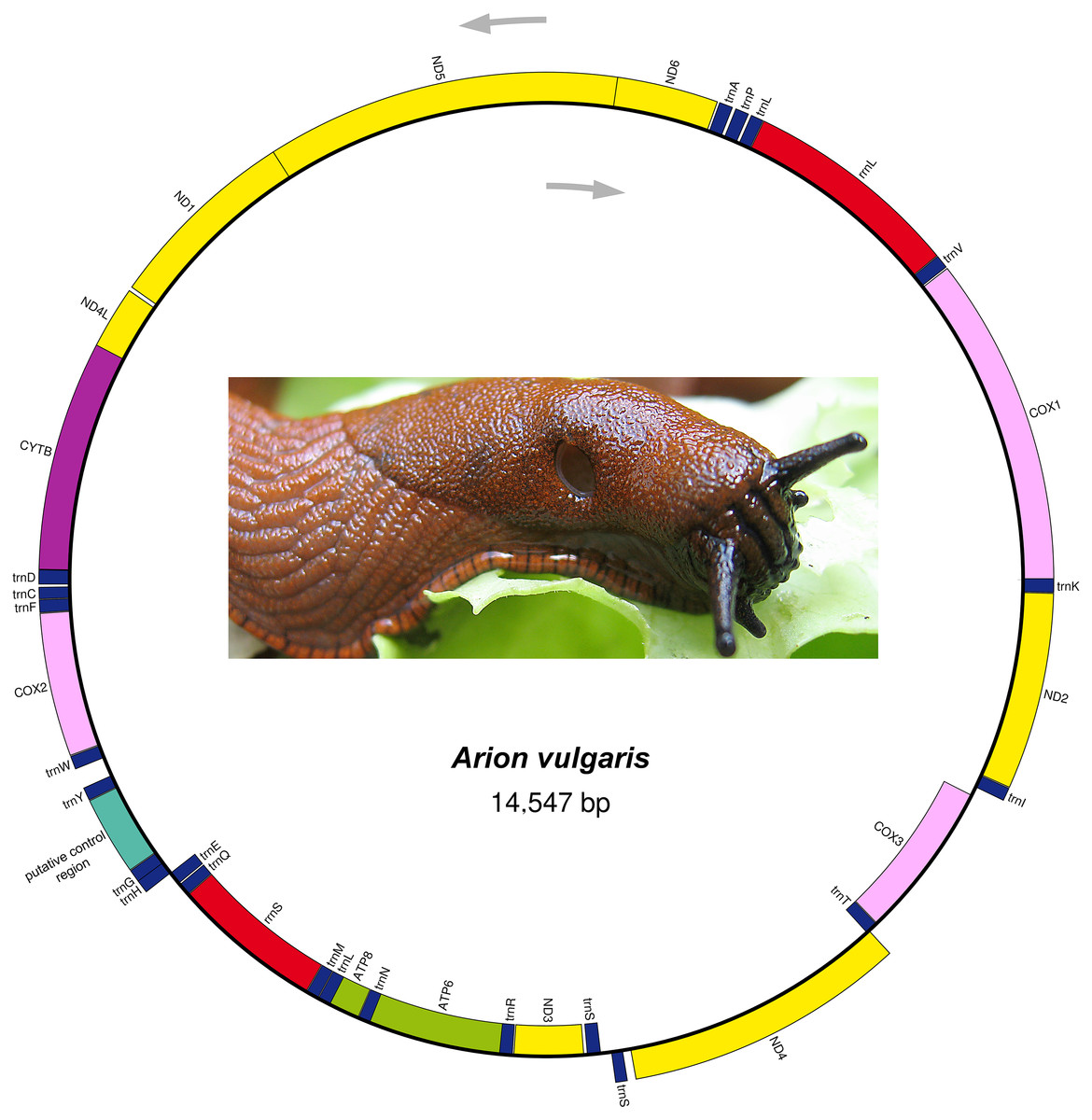
Mitogenome organization of Arion vulgaris
Furthermore, selection analyses were performed to investigate the role of different selective forces shaping stylommatophoran mitogenomes. Although purifying selection is the predominant selective force shaping stylommatophoran mitogenomes, six genes (ATP8, COX1, COX3, ND3, ND4 and ND6) detected by the branch-specific aBSREL approach and three genes (ATP8, CYTB and ND4L) detected by codon-based BEB, FUBAR and MEME approaches were exposed to diversifying selection. The positively selected substitutions at the mitochondrial PCGs of stylommatophoran species seems to be adaptive to environmental conditions and affecting mitochondrial ATP production or protection from reactive oxygen species effects. Comparative analysis of stylommatophoran mitogenome rearrangements using MLGO revealed conservatism in Stylommatophora; exceptions refer to potential apomorphies for several clades including rearranged orders of trnW-trnY and of trnE-trnQ-rrnS-trnM-trnL2-ATP8-trnN-ATP6-trnR clusters for the genus Arion. Generally, tRNA genes tend to be rearranged and tandem duplication random loss, transitions and inversions are the most basic mechanisms shaping stylommatophoran mitogenomes.